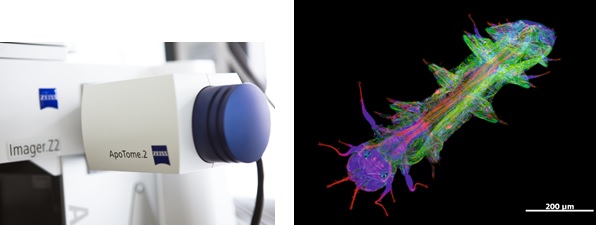
ApoTome – an innovative slider module for your Axio Imager, Observer or AxioZoom 16 fluorescence microscope that will change the way you view fluorescence microscopy.
The decisive step forward in performance for your lab. With the new ApoTome, you can expand the capacity of your digital imaging workstation in neuro, cell and developmental biology to exciting heights.
Significant advantages of the ApoTome system include:
• A performance leap in resolution
• Easy and seamless integration
• Minimal training necessary
• Spectacular results in minutes
The Principle
The imaging principle used with the ApoTome is referred to in the literature as “grid projection” or “structured illumination”. The fundamental theory was described in publications on interferometry as early as in the nineteen-sixties.
A grid of stripes of defined width is projected onto the focal plane of the objective and shifted laterally in three defined steps relative to the sample (see illustration). A CCD camera takes a picture in each grid position. The three “raw” images are combined into a resultant image by on-line computation. The resultant image is an optical section through the sample, from which all image slices originate. Free of artifacts, with out-of-focus information removed. This image has an improved signal-to-noise ratio, and approximately doubles resolution in axial (Z) direction. Making an optical section is a prerequisite for computing 3D visualizations of specimens.
Why is the resulting image obtained with structured illumination an optical section?
When we observe a specimen with a microscope, we can distinguish between two ranges in the axial (Z) dimension. All specimen details located within the depth-of-field range of the objective used are seen in sharp focus (the yellow area in the schematic illustration). Specimen details outside that range appear out of focus, i.e. blurred. In fluorescence microscopy, the out-of-focus portions of the image strongly interfere with the image information of the depth-of-field range.
The grid of stripes is imaged in the depth-of-field range of the objective. However, same as the specimen image, the grid image is in focus within a narrow depth range only. Below and above this in-focus range, the grid image gets out of focus and finally disappears (see illustration). If the grid is shifted to three different positions (see animation), the image data (gray levels) originating from out-of-focus ranges of the three raw images do not differ. As the data (gray levels) from outside the depth-of-field range are homogeneous, the algorithm used can remove the out-of-focus shares of the image.
Inside4D
The capability of ApoTome to make optical sections is a prerequisite for the acquisition of a stack of images and subsequent 3D reconstruction (3D rendering). For the reconstructions, you can use Inside4D, an optional AxioVision module with various rendering modes (shadow, transparency, surface or maximum projection). This module also allows you to process time series image stacks, and to create AVIs and Quicktime movies of reconstructions for integration into PowerPoint presentations, etc.
Download an APOTOME brochure.
Download an APOTOME MICROSCOPE ARTICLE.pdf brochure.